Our paper was accepted at CVPR2026 and selected as oral presentation!
Thrilled to share our accepted paper at CVPR2026 (Oral presentation)
Overview
Shuaike Shen, Ke Liu, Jiaqing Xie, Shangde Gao, Chunhua Shen, Ge Liu, Mireia Crispin-Ortuzar and Shangqi Gao. R2Seg: Training-Free OOD Medical Tumor Segmentation via Anatomical Reasoning and Statistical Rejection
Abstract. Foundation models for medical image segmentation struggle under out-of-distribution (OOD) shifts, often producing fragmented false positives on OOD tumors. We introduce R2-Seg, a training-free framework for robust OOD tumor segmentation that operates via a two-stage Reason-and-Reject process. First, the Reason step employs an LLM-guided anatomical reasoning planner to localize organ anchors and generate multi-scale ROIs. Second, the Reject step applies two-sample statistical testing to candidates generated by a frozen foundation model (BiomedParse) within these ROIs. This statistical rejection filter retains only candidates significantly different from normal tissue, effectively suppressing false positives. Our framework requires no parameter updates, making it compatible with zero-update test-time augmentation and avoiding catastrophic forgetting. On multi-center and multi-modal tumor segmentation benchmarks, R2Seg substantially improves Dice, specificity, and sensitivity over strong baselines and the original foundation models.
OOD test-time adaptation
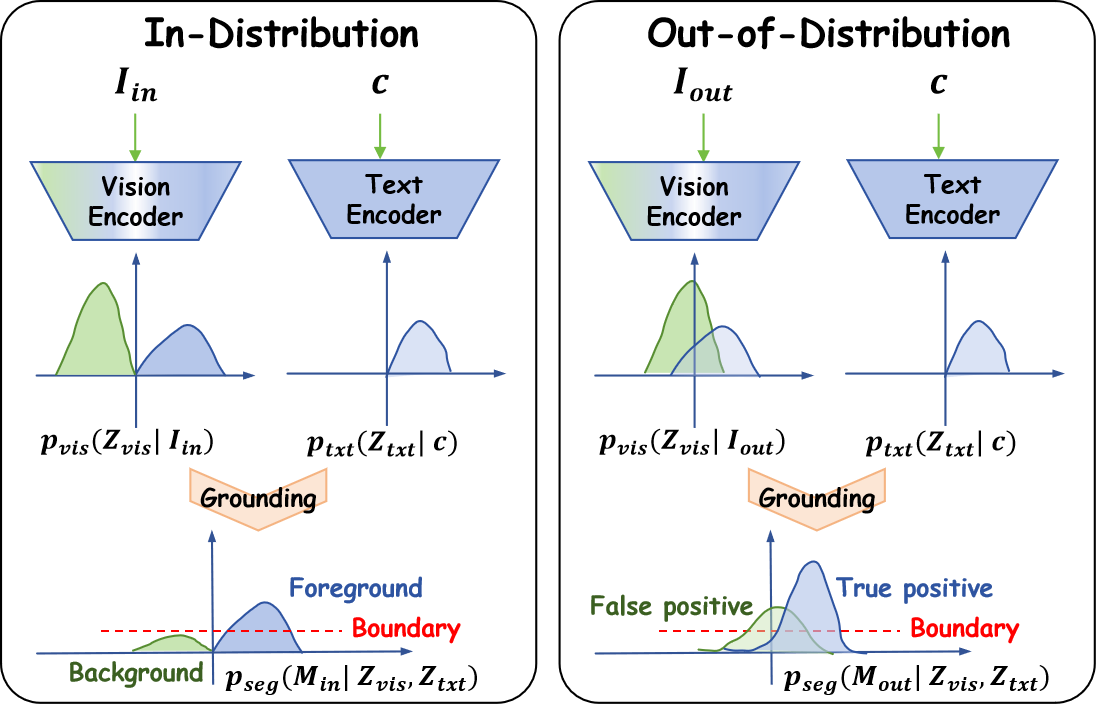
When vision embeddings are well separated, the model can distinguish foreground from background by aligning text embeddings with a single decision boundary (left). In medical imaging, however, protocols vary across scanners, tumor sites, and modalities. Vision embeddings for OOD samples become poorly separated, biasing the decision boundary so that background structures are misclassified as tumors — leading to high false-positive rates and potentially harmful over-diagnosis (right).
Architecture of R2Seg